 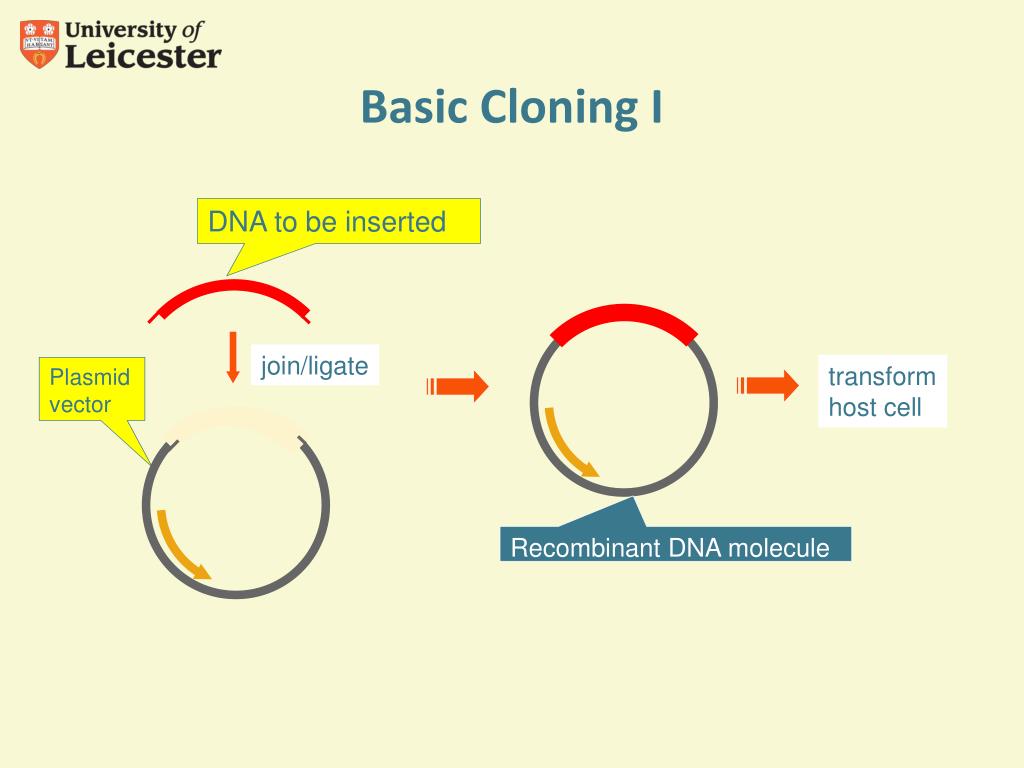 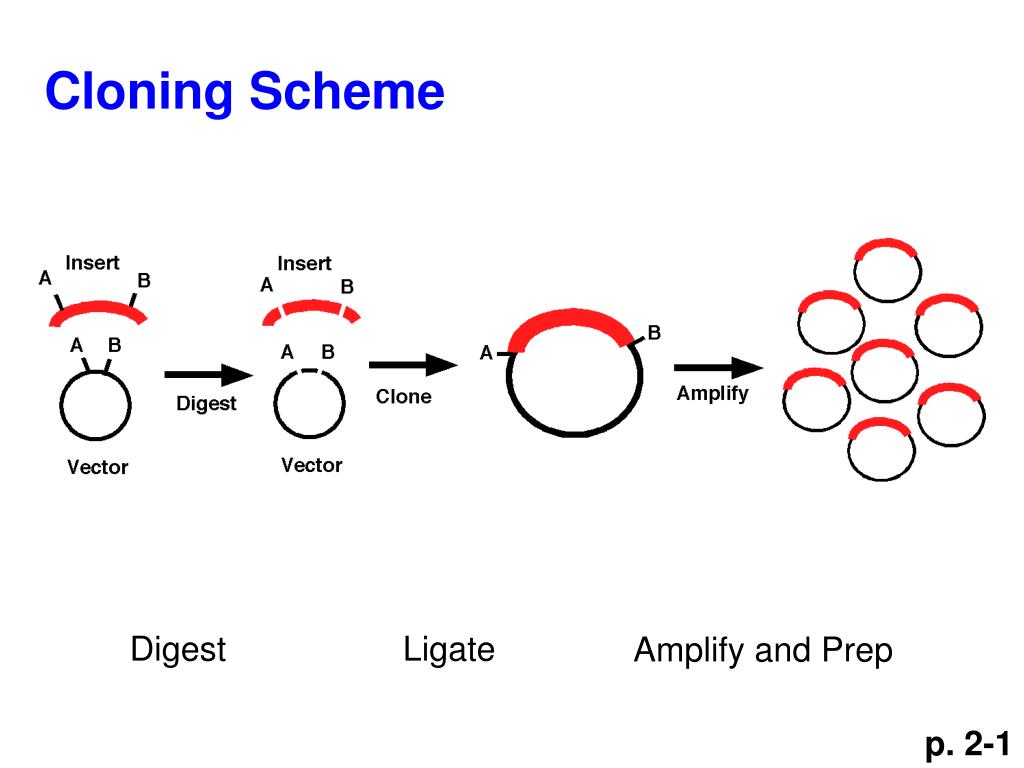
Briefly, the sequences to be joined need to be flanked by mutually exclusive variants of the attachment sites B and P or L and R. However, the 34 basepairs (bp) of the loxP sites lead to long cloning scars and the number of elements that can be combined is limited to two.Īnother very popular recombination based cloning method is Invitrogen’s Gateway® technology, which exploits the lambda phage’s integration and excision mechanism. Here, a gene of interest was cloned in the so-called pUNI vector and then transferred into a pHOST vector providing the regulatory sequences for expression, which allowed standardized and quick shuffling between a collection of entry and destination vectors. One of the first ligation independent cloning methods was the univector plasmid fusion system, which is based on Cre/ loxP-mediated recombination. With the advent of high throughput approaches and widespread use of transgenic models to test gene function in vivo, classical restriction based cloning rapidly became a major limitation and alternative technologies began to emerge. To overcome these limitations, researchers had to devise complicated cloning strategies to assemble plasmids, which involved many steps and required a large number of diverse restriction enzymes. Furthermore, many of these recognition sites occur fairly frequently in a given piece of DNA, making the assembly of long constructs even more difficult because of the lack of unique restriction targets. However, since most recognition sites used in a digestion-ligation cycle remain in the construct, the corresponding enzymes cannot be used for adding further DNA fragments in subsequent steps. Thus, in case more than four DNA elements have to be assembled, success-rates drop significantly and additional rounds of cloning may be necessary. While this approach is generally successful, there are certain limitations, especially when it comes to the assembly of complex plasmids from multiple elements, since with increasing numbers of DNA fragments the ligation reaction becomes less and less efficient. The DNA fragments to be combined are first excised from their precursor molecules via the endonucleases and then in a separate reaction re-assembled by the ligase, usually after spontaneous annealing of complementary single-stranded overhangs created during the endonuclease cut. The system is cheap and reliable and when combined with a library of modules considerably speeds up cloning and transgene stacking for plant transformation.Įver since the first construction of a recombinant plasmid, , genetic engineering and molecular cloning mostly rely on the use of type II restriction endonucleases and DNA ligases. 
GreenGate cloning is simple and efficient since it uses only one type IIS restriction endonuclease, depends on only six types of insert modules (plant promoter, N-terminal tag, coding sequence, C-terminal tag, plant terminator and plant resistance cassette), but at the same time allows assembling several expression cassettes in one binary destination vector from a collection of pre-cloned building blocks. Here we introduce our GreenGate system for rapidly assembling plant transformation constructs, which is based on the Golden Gate method. Several alternative cloning systems have been developed in recent years to overcome these issues, including the type IIS enzyme based Golden Gate technique. Especially in case of long inserts and applications requiring high-throughput, this approach is limited by the number of available unique restriction sites and the need for designing individual cloning strategies for each project. Traditionally it is based on a multitude of type II restriction endonucleases and T4 DNA ligase. Building expression constructs for transgenesis is one of the fundamental day-to-day tasks in modern biology. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
